Genetic evidence of ancient migration routes along the Atlantic Coast of South America are expected.
Strong Denisovan ancestry and an Australasian signal are not, unless the signals have shallow time depth. The Panamanian sample newly identified as having such a signal is indeed recent, less than two hundred and fifty years post-Columbian and is not found in seven other ancient DNA samples at the site in the same study.
An increasing body of archaeological and genomic evidence has hinted to a complex settlement process of the Americas. This is especially true for South America, where unexpected ancestral signals have raised perplexing scenarios for the early migrations into different regions of the continent.
Here we present ancient genomes from the archaeologically rich Northeast Brazil and compare them to ancient and present-day genomic data.
We find a distinct relationship between ancient genomes from Northeast Brazil, Lagoa Santa, Uruguay and Panama, representing evidence for ancient migration routes along South America’s Atlantic coast.
To further add to the existing complexity, we also detect greater Denisovan than Neanderthal ancestry in ancient Uruguay and Panama individuals. Moreover, we find a strong Australasian signal in an ancient genome from Panama.
This work sheds light on the deep demographic history of eastern South America and presents a starting point for future fine-scale investigations on the regional level.
Andre Luiz Campelo dos Santos, et al., "Genomic evidence for ancient migration routes along South America’s Atlantic coast" bioRxiv (July 1, 2022) (Supplemental Materials here).
The body text states:
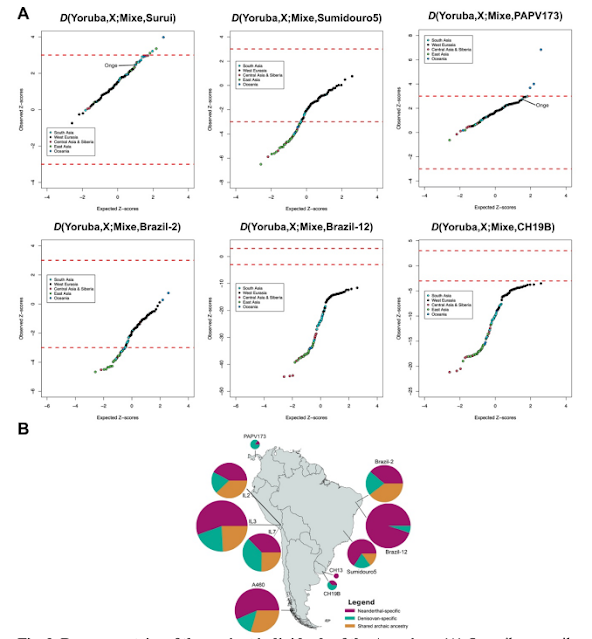
Fig. 2. Deep ancestries of the ancient individuals of the Americas. (A) Quantile–quantile plots of the Z-scores for the D-statistic test for each ancient sample, a non-American, non-African population ‘X’ from Simons Genome Diversity Project (SGDP), and the Mixe, compared to the expected ranked quantiles for the same number of normally distributed values. Each data point represents a distinct population from SGDP. Dashed red lines represent significance thresholds. (B) Archaic ancestry in ancient South America and Panama. Pie chart radius reflects the proportion of shared archaic loci in the individual.
Overall, our results show a strong genomic relationship among Brazil-2, CH19B, Sumidouro5 and PAPV173 (Fig. 1, B to D, and Fig. 4). Apart from the occurrence of mass burials in the sites that yielded these samples, there is no other evidence in the archaeological record that indicate shared cultural features between them. It is also important to note that Sumidouro5 is ~9,000 years older than the other three mentioned individuals, enough time for expected and noticeable cultural divergence. Moreover, Brazil285, CH19B and PAPV173, though more similar in age, were located thousands of kilometers apart from each other. Therefore, cultural differentiation is also expected among them. On the other hand, our results further corroborate previously reported evolutionary relationships between PAPV173 and CH19B by providing evidence of a distinct genomic affinity involving archaic human ancestry (Fig. 2B and fig. S4).A strong signal of Australasian ancestry, previously reported only for the Lagoa Santa individual and present-day Surui, was also observed for the previously published PAPV173, from Panama (Fig. 2A). The Piapoco, Surui and Karitiana, however, harbor high affinities with Brazil-2 (Figs. 3 and 4A), and thus may have received contributions coming from Central America (in the form of the Australasian signal, in a north-to-south directionality) and South America’s Atlantic coast (in a south-to-north directionality).
The Panamanian sample is a man from the late 14th or early 15th century on Panama's Pacific Coast and is reported in Capodiferro MR, et al. "Archaeogenomic distinctiveness of the Isthmo-Colombian area." 184 Cell 1706-1723.e24 (2021) (doi:10.1016/j.cell.2021.02.040)
The introduction to that paper states:
Archaeological and genetic evidence suggests that the peopling of sub-Arctic America started from Beringia before, during, and immediately after late Glacial times (Achilli et al., 2018; Ardelean et al., 2020; Becerra-Valdivia and Higham, 2020; Braje et al., 2017; Skoglund and Reich, 2016; Waters, 2019; Yu et al., 2020). Initial settlement attempts were followed by a more widespread peopling that reached southern South America as early as ∼15 thousand years ago (kya) (Dillehay et al., 2017).
Recent studies of ancient and modern genomes describe a complex scenario prior to European contact with multiple migrations from Beringia, as initially suggested by mitochondrial DNA (mtDNA) data (Achilli et al., 2013; Brandini et al., 2018; Gómez-Carballa et al., 2018; Llamas et al., 2016; Perego et al., 2009; Perego et al., 2010; Tamm et al., 2007) as well as demographic spreads and admixture events along the two continents (Flegontov et al., 2019; Moreno-Mayar et al., 2018; Posth et al., 2018; Scheib et al., 2018; Schroeder et al., 2018).
The great majority of ancestries in early Native Americans (NAs, here used to indicate Indigenous groups) derive from an ancestral Beringian population(s) that differentiated sometime between ∼22 and ∼18 kya and likely exhibited genetic sub-structure that may explain the initial late Glacial migration(s) as well as the spread of the so-called UPopA (unknown population in the Americas) whose legacy reappears in Central America ∼8.7 kya, leaving signs in the gene pool of the Mixe (Moreno-Mayar et al., 2018).
In unglaciated eastern Beringia/northern North America, the first peoples split into two branches called Northern NA (NNA, or ANC-B) and Southern NA (SNA, or ANC-A). The most ancient representatives of SNA are individuals who were living on both sides of the Rocky Mountains more than 10 kya: the Clovis-associated Anzick-1 and the Spirit Cave individuals associated with Western Stemmed technology.
Ancient individuals carrying SNA ancestries crossed the Panama land bridge and entered South America. Their fast spread along the southern continent is evidenced by the earliest archaeological human presence in the Southern Cone at 14.6 kya and by ancient human genomes dating more than 9 kya on both sides of the continent: at Cuncaicha (Peru) and Los Rieles (Chile) on the Pacific and Lapa do Santo and Lagoa Santa (Brazil) on the Atlantic.
Another UPop (UPopY) with Australasian ancestry may have contributed to the early peopling of South America as recognized in one sample from the Lagoa Santa site and in some Amazonian groups that experienced isolation events (e.g., Surui and Karitiana) (Moreno-Mayar et al., 2018; Skoglund et al., 2015).However, the demographic dynamics underlying many of these events, before and after European contact, are still uncharacterized, especially at the regional level (Fernandes et al., 2020; Lindo et al., 2017; Nakatsuka et al., 2020; Nägele et al., 2020).
The Panamanian isthmus lies between the Atlantic and Pacific oceans and connects the two American continents. It was the only land bridge during the initial peopling of South America and has remained a crossroads of goods, technologies, ideas, and peoples throughout history, including more recent colonial times (Cooke, 2005; Cooke et al., 2019; Hernández Mora et al., 2021). In light of Panama’s geographic location, the archaeogenomic study of its past can reveal its demographic history, including movements between North and South America.






4 comments:
The preference for the Onge over the Papuan sample suggests a non-Australasian source of that ancestry, no?
Fascinating to see this New America onion being peeled. So much history here.
@Ryan
Honestly, the analysis in this particular pair of papers could be better. I'm not sure what is going on. A paper in Nature suggested that it is a basal East Eurasian ancestry, possibly from NE Asia.
I remain pretty comfortable that the time depth is just not that great, however, probably in the last 1200-600 years or so.
What do you mean you're not sure what it's going on? The Z score for Sumidouro5 is significant.
And the time depth is more than that. Sumidouro5 is 10kya.
Sorry but your model is out.
Post a Comment